Interpretacion: The CISH hybridization signal for a single copy of HER2 gene appears as a distinctive mark, as a dark green dot while the signal from the centromeric region of chromosome 17 as a punctate signal of bright red color, which can be distinguished clearly by the background counterstaining with hematoxylin. The visualization of the signals must be made with an objective of at least 40x. No filter lens to enhance the contrast should be used as this may distort the color signal. For brightly colored signals is desirable to increase the diaphragm aperture. Also make sure to focus up and down when doing a core assessment as the red and green signals may be located in different planes.Prior to quantify the signal, the tissue section / extended cells must be scanned with an objective of 10x or 20x to detect any possible intratumoral heterogeneity. If heterogeneity exists, for the evaluation should be chosen a
representative area for the amplification status of the tumor. For the numerical quantification of the signals, necrotic
areas, overlapped nuclei and weak signal intensity should be avoided.
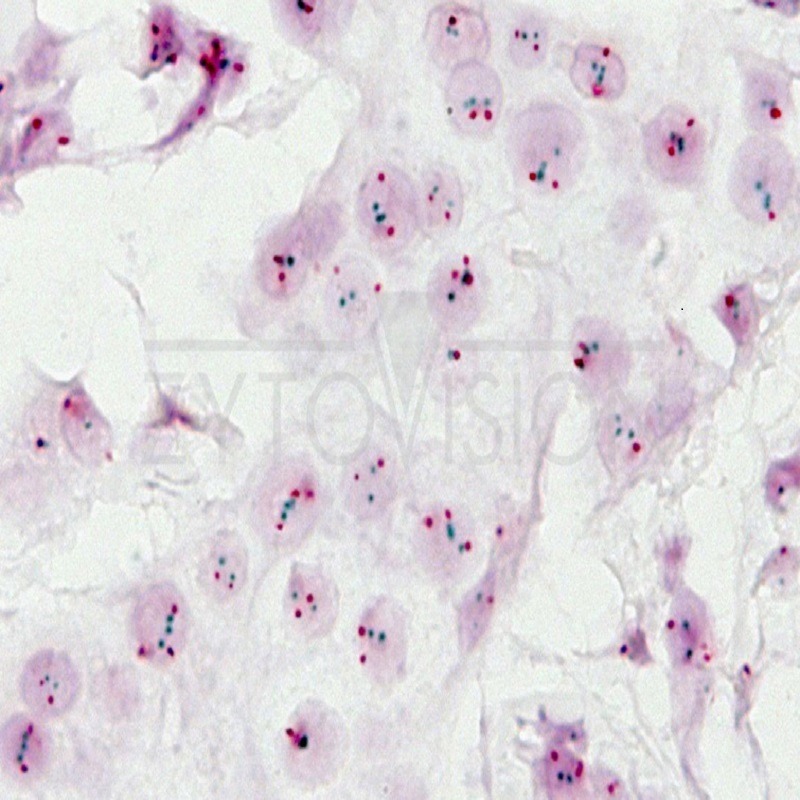
In normal diploid nuclei without amplification of the gene, even in normal cells or other accompanying cells, in each
nucleus two punctate green and red signals with smooth contour are visible (see Fig. 1). Due to mitosis, additional
signals might be seen even in a small percentage of non-neoplastic cells. Occasionally, because the signals might be
located above or below the plane of the section, cells with no signal could be seen.
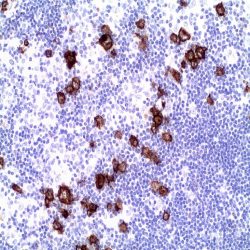
If aneuploidy of chromosome 17 (see fig. 2) multiple additional red signals will be visible.
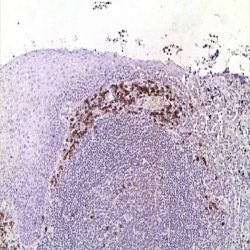
If HER2 gene amplification (see Fig. 3) the specific green HER2 gene signals are visible as multiple points or small groups
of irregular shape. As a reference for the green signal, normal cells present in the same preparation should only be
used. In addition, red centromere signals will be clearly visible.
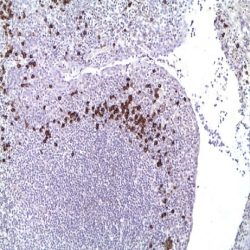
In case of high gene amplification many green dots or large groups will be visible in the nuclei (see fig.4). As a reference
for the green signal, normal cells present in the same preparation should only be used. Red centromeric signals may be
superimposed and may not be apparent or visible.
However for the correct interpretation of the results is recommended to follow the latest guidelines for effective
assessment for each type of tissue used in the published literature.
Limitation Of Reactive: The use of frozen tissue has not been evaluated.
Sample Types: Sections of 4 microns thick mounted on special slides for immunohistochemistry and obtained from paraffin-embedded
tissues, preferably fixed in buffered formalin.
Principle Of Analytical Method (in situ hybridization): In situ hybridization (ISH) allows the detection of specific sequences of DNA or RNA in histological and cytological
samples without losing morphological details. The ISH technique is based on hybridization between a DNA or RNA sequence specifically labelled (probe) and a sequence of DNA or RNA present in the sample. If there is any complementarity between the sequences the hybridization occurs, which results in a hybrid product. Hybrids can be easily viewed by a chromogenic immunohistochemical staining procedure (CISH) directed toward specific marker of the probe employed. The ISH technique is highly sensitive, specific and easy to perform, and in addition, contains no radioactive products. The reagents supplied in this kit are perfectly matched and therefore, the kit is an easy to use tool to perform CISH.

ماندان –
شما کانال تلگرام هم دارین؟
مدیریت –
سلام. بله دوست عزیزم. در تلگرام جستجو کنید : samatashkhis